Project Details
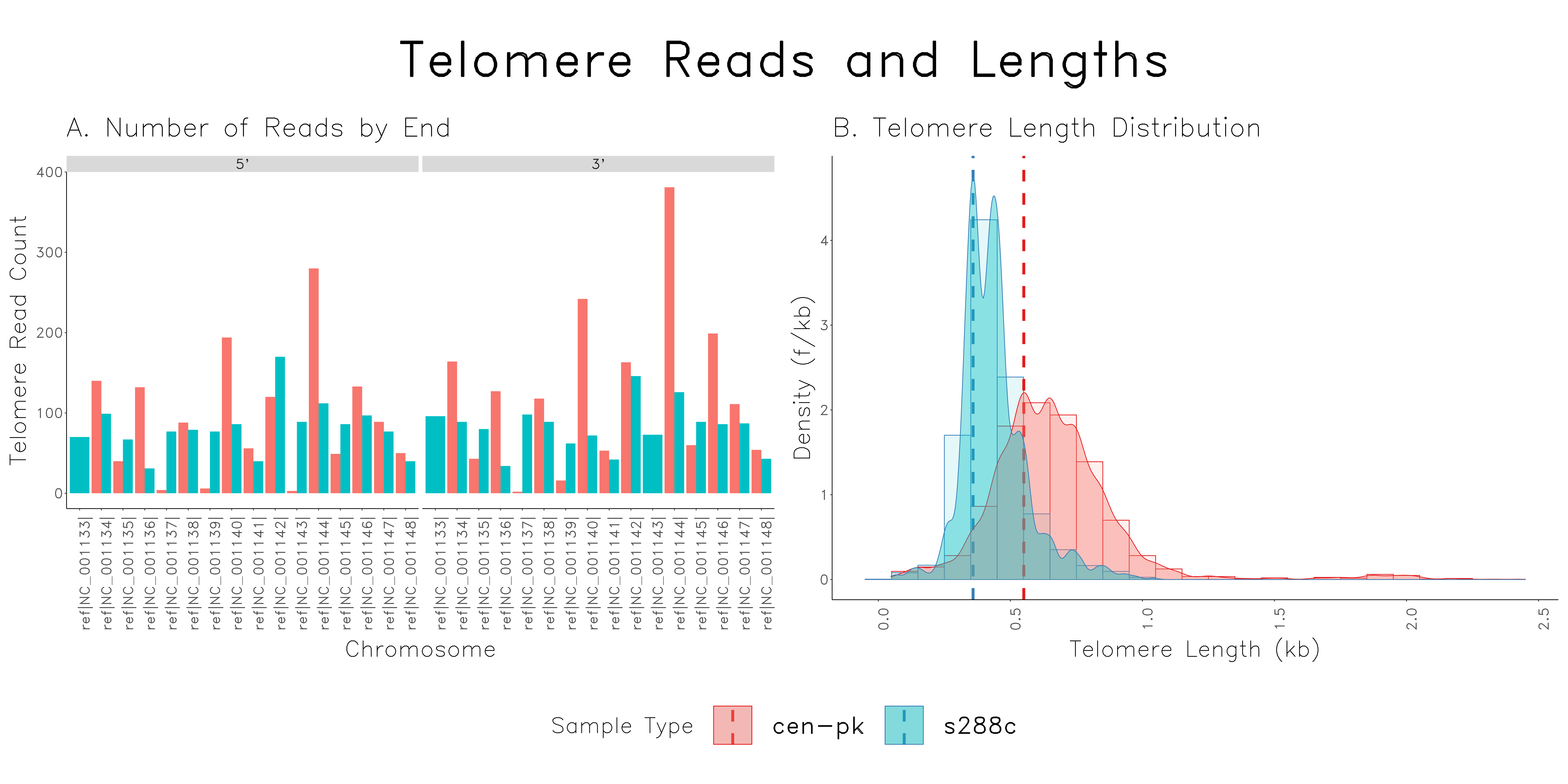
I created TLD as a solution for telomere length analysis in long-read sequencing data. TLD is a package that uses a combination of R and Python to analyze telomere end chromosome data and produce visualizations. The package was designed with reproducibility in mind; thus, is it containerized and available on DockerHub and GitHub. The application was designed specifically for analyzing telomere length in Plasomodium falciparum samples which had been irradiated. The purpose of this study was to determine the effects of radiation on telomere length in the parasite and therefore, inform our understanding of telomere biology. The study was able to replicate a finding in the NASA Twin Study which found that the astronaut in space had longer telomeres which recovered to normal lengths after being on the ground for two months. The study was published in iScience and is available here. The above image is the result of analyzing real data from a Saccharomyces cerevisiae sample.